Abstract
MicroRNAs (miRNAs) control the expression of protein-coding genes in normal hematopoietic cells and, consequently, aberrant expression may contribute to leukemogenesis. To identify miRNAs relevant to pediatric acute lymphoblastic leukemia (ALL), we cloned 105 known and 8 new miRNA genes expressed in patients’ leukemia cells. Instead of known miRNA genes, new miRNA genes were not evolutionarily conserved. Quantification of 19 selected miRNA genes revealed an aberrant expression in ALL as compared with normal CD34+ cells (P⩽0.02); both upregulated (14/19) and downregulated (5/19) expressions were observed. Eight miRNAs were differentially expressed between MLL and non-MLL precursor B-ALL cases (P<0.05). Most remarkably, miR-708 was 250- up to 6500-fold higher expressed in 57 TEL-AML1, BCR-ABL, E2A-PBX1, hyperdiploid and B-other cases than in 20 MLL-rearranged and 15 T-ALL cases (0.0001< P<0.01), whereas the expression of miR-196b was 500-fold higher in MLL-rearranged and 800-fold higher in 5 of 15 T-ALL cases as compared with the expression level in the remaining precursor B-ALL cases (P<0.001). The expression did not correlate with the maturation status of leukemia cells based on immunoglobulin and T-cell receptor rearrangements, immunophenotype or MLL-fusion partner. In conclusion, we identified new miRNA genes and showed that miRNA expression profiles are ALL subtype-specific rather than linked to the differentiation stadium associated with these subtypes.
Similar content being viewed by others
Introduction
Recently, a new class of ∼22-nucleotides endogenous RNAs, called microRNAs (miRNAs), has been discovered. Despite their name, these small RNAs have a major biological function by affecting the translation of proteins encoded by many genes.1, 2 Although previous and current cancer research mainly focused on protein-coding genes, the function of miRNAs in gene regulation emphasizes the need to address the contribution of non-coding small RNAs in cancer cell physiology. miRNAs are encoded by at least 710 genes in humans, and are estimated to represent 1–5% of all predicted human genes.3, 4 They form one strand of the miRNA:miRNA* duplex that is processed from a larger stem-loop precursor.5, 6, 7, 8, 9 Only the mature miRNA strand binds to its (imperfect) complementary messenger RNAs (mRNAs), resulting in the translational repression of the targeted mRNA.10, 11, 12, 13 Currently characterized miRNAs inhibit translation of proteins that play important roles in fundamental biological processes, such as proliferation and differentiation,2, 14, 15 processes that are also affected in cancer cells. The expression profiles of miRNAs have been shown to be altered in various tumors including lung cancer,16 colon cancer17 and different types of leukemia, for example chronic lymphocytic leukemia18 and acute myeloid leukemia (AML).19, 20 Oncogenic activity of miRNAs was shown by the fact that introduction of the miR-17-92 cluster, which is upregulated in malignant lymphomas, accelerated c-myc-induced lymphomagenesis in mice.21
MiRNA expression levels may be used in the classification of cancer. Lu et al. showed that poorly differentiated tumors could be more accurately identified using miRNA expression profiles compared to mRNA expression profiles.36 More recently, miRNA signatures were also shown to classify different cytogenetic entities of adult AML, and aberrant miRNA expression was shown to be linked to the prognosis of adult AML.19, 20
In contrast to several solid tumors, lymphomas, chronic lymphocytic leukemia and AML, the importance of miRNAs in acute lymphoblastic leukemia (ALL) is yet largely unknown. ALL represents a heterogeneous disease characterized by various underlying genetic abnormalities. Intensive combination chemotherapy schedules have resulted in a 5-year event-free survival of ∼80% in children, whereas the treatment of adult ALL is much less successful resulting in ∼40% 5-year event-free survival.22 Genetically, different subtypes of ALL have different clinical outcomes. For example, infants (children <1 year) with ALL cells bearing a rearrangement of the MLL gene have a highly unfavorable 4-year prognosis of <40%.22 It is not unlikely that miRNAs also play a role in ALL, as its subtypes are characterized by differentiation arrests at various stages in normal B- and T-cell development.
Present miRNA expression array-detection techniques are based on miRNA genes that have been published in the Sanger database. Consequently, one may miss yet-unknown miRNAs that are especially expressed in leukemic cells. Hence, we first systematically cloned miRNA genes that are expressed in leukemic cells of pediatric ALL patients with a poor prognostic MLL gene rearrangement and a prognostically more favorable precursor B-ALL subtype (B-other) to identify ‘leukemia-specific miRNAs.’ In total, 105 known and 8 new miRNA genes were identified in these two subtypes. Quantification of miRNA expression by stem-loop real-time PCR analysis revealed that the expression of both known and newly identified miRNA genes differed between genetically and prognostically different subgroups of ALL and between these subtypes and normal CD34+ progenitor cells. Our data show that the current number of miRNA genes relevant to ALL is underestimated and that systematic miRNA gene cloning and subsequent miRNA expression analysis reveals ALL subtype-specific miRNA profiles. Our data also indicate that these subtype-related differences in miRNA expression levels cannot be explained by differences in the maturation status of individual ALL cases as judged on immunoglobulin and T-cell receptor rearrangement patterns and CD marker expression. Hence, the miRNA expression pattern seems more subtype-specific than B- or T-cell differentiation status specific in pediatric ALL.
Materials and methods
Patient samples
After obtaining informed consent, peripheral blood or bone marrow samples were obtained from children with ALL at primary diagnosis. MLL gene rearranged precursor B-ALL samples were collected from newly diagnosed infants (<1 years of age) who participated in the Interfant study. All samples were screened for (specific) MLL gene rearrangements by reverse transcriptase-PCR (RT-PCR) and fluorescent in situ hybridization. A total of 20 MLL-rearranged samples included in this study are positive for t(4;11), n=8; t(11;19), n=8; t(9;11) n=3 and t(1;11), n=1. Fifteen T-ALL samples and 57 CD19+ precursor B-ALL samples were obtained from the Cooperative Study Group for Childhood Acute Lymphoblastic Leukemia study (COALL; Hamburg, Germany) and the Erasmus MC-Sophia Children's Hospital (Rotterdam, The Netherlands). All non-infant precursor B-ALL samples were negative for MLL translocation and were genetically characterized by the presence of hyperdiploidy (more than 50 chromosomes, n=10), the TEL-AML1 (n=10), BCR-ABL (n=10) and E2A-PBX (n=8) translocations. The remaining 19 precursor B-ALL samples were negative for hyperdiploidy, MLL, TEL-AML1, BCR-ABL and E2A-PBX translocations (B-other). CD34+ control samples were obtained from granulocyte colony-stimulating factor-mobilized peripheral blood stem cell harvests from two children with a brain tumor after informed consent.
Isolation of RNA and DNA out of leukemic cells
Mononuclear cells were isolated out of primary bone marrow and peripheral blood samples as described earlier.23. Non-malignant cells were removed using immunomagnetic beads. All processed leukemia samples contained >90% blast cells, as determined on May–Grünwald–Giemsa (Merck, Darmstadt, Germany)-stained cytospin preparations of isolated cells. CD34+ control cells were enriched using magnetic beads (Miltenyi Biotec, Bergisch Gladbach, Germany) specific for CD34. Purity (>90%) was assessed using flow cytometry. A minimum of 5 × 106 cells was lysed in TRIzol reagent (Invitrogen, Breda, The Netherlands) to extract total RNA.23 Quality of RNA was examined by the 2100 bioanalyzer (Agilent Technologies, Amstelveen, The Netherlands).
Analysis of the maturation status of MLL-rearranged precursor B-ALL and T-ALL cases
Ig/TCR rearrangement patterns of MLL-translocated ALL samples were analyzed by PCR as described earlier.24. Three stages of B-cell maturation were defined, namely immature (no or only incomplete IGH (DH-JH) rearrangements and no detectable TCR gene rearrangements), intermediate (incomplete (DH-JH) or complete (VH-JH) IGH rearrangements and/or incomplete (Dδ2-Dδ3 or Vδ2-Dδ3) TCRD rearrangements; no other Ig/TCR gene rearrangements detectable) and mature (Vδ2-Jα rearrangement and/or IGK rearrangement and/or IGL rearrangement and/or TCRG rearrangement and/or TCRB rearrangement). The maturation status of T-ALL samples was based on the expression of CD1 and surface-membrane-bound CD3 as determined by flow cytometry (SmCD3−/CD1−=immature, SmCD3−/CD1+=intermediate and SmCD3+=mature).
Direct cloning of miRNAs
We used a slightly modified version of the direct cloning as described earlier by Lau et al.25 Briefly, 1 pmol of 32P-labeled 23-mer Carrier Oligo (5′-UGUCAGUUUGUUAAUUAACCCAA-3′) was spiked into a minimum of 1.5 μg total RNA extracted from purified leukemic cells. Subsequently, RNA size fractionation was performed on a 15% polyacrylamide 8 M urea gel (National Diagnostics, Atlanta, Georgia, USA) and 18–26 nucleotides small RNAs were isolated using the radiolabeled oligo as a reference. The 18–26 nucleotides-fractionated small RNAs were first ligated to a 3′-adaptor oligonucleotide (5′-pre-adenylated) without the presence of ATP, and then to a 5′-adaptor oligonucleotide (not adenylated) in the presence of ATP. Final ligation products were amplified by RT-PCR, after which the PCR products were digested with PacI restriction enzyme (New England Biolabs (NEB)), Ipswich, MA, USA) to eliminate carrier oligo products. After precipitation, PacI-digested PCR products were purified on a 15% non-denaturing polyacrylamide gel (National Diagnostics) and, according to their size, isolated from the gel and further amplified by 10–20 PCR cycles. PCR products were then digested with BanI restriction enzyme (NEB) and, if necessary, once more with PacI restriction enzyme. This was followed by concatemerization of BanI restricted products and subsequent ligation into the pCR 2.1-TOPO vector (Invitrogen). Finally, the sequence of small RNA libraries was analyzed (Macrogen, Seoul, Korea), and the data were bioinformatically analyzed to identify which miRNAs were expressed.
Bioinformatic analysis of small cloned RNAs
Cloned small RNA sequences were mapped to human and mouse genomes. Candidate precursor miRNA sequences were selected and computationally folded with the Vienna RNA Package.26 Folded precursors were identified as miRNA candidates by applying a set of parameters derived from known miRNA genes as shown by Supplementary Figure S1. The following parameters were applied to human miRNA candidates: a loop length of 6–35 bp; a 15–36 bp length of the miRNA* sequence, which is defined as the (imperfect) complementary strand of the miRNA within the folded precursor structure; a percentage of 55–100% of the miRNA that binds to the miRNA* sequence with 100% complementarity; and finally an energy from the precursor miRNA sequences between −42.5 and −6.8 kcal/mol. In case of mouse miRNA precursors, a loop length of 7–35 bp, a miRNA* length of 15–28 bp, a pairing percentage between 67 and 100% and an energy between −46.4 and −14.2 kcal/mol were applied.
Stem-loop RT-PCR
MiRNA expression was measured by stem-loop RT-PCR using primers and probes of the TaqMan MicroRNA assay for yet-known miRNAs or newly developed primers and probes for newly identified miRNA candidates (Applied Biosystems, Foster City, CA, USA).27 Endogenous small nucleolar RNA 1 (snoRNA 1, 5′-AUUUGCUAUCUGAGAGAUGGUGAUGACAUUUUAAACCACCAAGAUCGCUGAUGCA-3′) or spiked synthetic miR-181c was used as a reference for RNA input. In the latter case, synthetic miRNA-181c (Applied Biosystems) was added to total RNA before reverse transcription. Real-time PCR was performed in triplicate on an Applied Biosystems 7900HT RT-PCR system.
Statistical analysis
The Mann–Whitney U-test was used to compare expression levels of miRNAs between two groups. Fisher's exact test was used for frequency variables. Differences were considered statistically significant when P<0.05, two-sided.
Results
Identification of new miRNAs in two subtypes of pediatric ALL
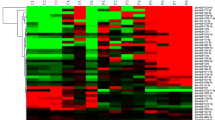
A total of 1128 and 1080 small RNA (sRNA) sequences were cloned from the leukemic cells of an MLL-rearranged patient carrying a t(1;11) translocation and a precursor B-ALL patient negative for all known genetic subtypes in ALL (referred to as B-other), respectively (Figure 1a). Sequences of cloned sRNAs were mapped to the human genome and sequences encoding protein-coding mRNAs were excluded.28 Precursor miRNA candidates were then extracted, folded to create the initial miRNA duplex and evaluated on the basis of features derived from a set of known miRNAs present in the Sanger database as described in the section Materials and methods.4 Ninety-five and 88 miRNA genes were found to be expressed in the leukemic cells of the MLL-rearranged and B-other cases, respectively. A number of these genes were present in both patients, resulting in the identification of a total of 113 unique miRNA genes (Figure1a and Table 1 and Supplementary Table S1). Of these, eight genes (7%) are newly discovered miRNA genes encoding eight new mature miRNAs not reported earlier in miRBase v11.0.4 The 105 known genes (93%) encode 86 mature miRNAs and 15 miRNA* forms.
(a) Identification of miRNA genes expressed in leukemic cells of MLL-rearranged and other precursor B-ALL patients by direct cloning of small RNAs and subsequent bioinformatic analysis (see Materials and methods). (b) Evolutionary conservation of miRNAs that are expressed in ALL. Bars represent the percentage of miRNAs that share homology with mouse miRNAs (conserved; black) and of miRNAs without mouse homology (non-conserved; white). Conservation is shown for known mature and known star forms as well as for newly identified mature miRNAs.
Table 1 and Supplementary Table S1 show the clone count within the constructed libraries for newly cloned and known miRNAs. Newly identified miRNAs were cloned with a lower frequency than that of the majority of known miRNAs (P<0.001). For example, the maximum total clone count for newly found miRNAs is 4 (hsa-miR-1979), whereas the maximum clone count for known miRNAs is 667 (miR-142-3p). Another difference between newly found miRNAs and yet-known miRNAs is the fact that the sequences of the newly identified miRNA genes are not conserved between humans and mice. Ninety-two percent (79/86) and 80% (12/15) of the known mature miRNAs and miRNA*s share homology with mouse miRNAs, respectively (Figure 1b).
MiRNA expression profiles differ between MLL-rearranged and other precursor B-ALL cases
As the total clone count obtained by the summation of clones containing identical DNA sequences is not an accurate quantification method, miRNA expression levels were measured by stem-loop RT-PCR.27 Initially, the relative expression levels to spiked miR-181c of 8 novel and 86 known miRNAs/miRNA* forms were analyzed in the same samples that were used for direct miRNA cloning. MiRNA levels were compared with the level in CD34+ normal cells (Supplementary Tables S2 and S3). Supplementary Table S2 shows that despite their low clone frequency within the library (Table 1), the expression of newly identified miRNAs can be detected using the stem-loop RT-PCR technique. For example, hsa-miR-1975 was detected by just a single clone count, but this miRNA is 3.5-fold higher expressed in precursor B-ALL as compared with CD34+ using quantitative stem-loop RT-PCR.
Out of the total of 94 validated miRNAs, we selected a set of highly differentially regulated miRNA candidates for which quantitative stem-loop RT-PCR primers/probes were available (Supplementary Tables S2 and S3). These miRNAs were validated in a larger group of 16 MLL-rearranged and 19 other precursor B-ALL patients being negative for MLL, TEL-AML1, BCR-ABL, E2A-PBX and hyperdiploidy (B-other). In addition, five miRNAs were included that are located near the 11q23 chromosomal region involved in the MLL translocation (that is, miR-100, miR-34b and miR-181c) or in the function of MLL-1 protein (that is, miR-142-5p and miR-142-3p).29 As shown in Table 2, all 19 miRNAs are significantly differentially expressed in B-other ALL versus normal CD34+ blood cells (P⩽0.001), and 18 out of 19 miRNAs show a significant differential expression in MLL-rearranged ALL versus normal CD34+ cells (P⩽0.02). Expressions of 14 out of 19 miRNAs (miR-128a, miR-142, miR-150, miR-181, miR-30e-5p, miR-193, miR-34b, miR-365, miR-582, miR-708) were median 1.4- to 2534-fold upregulated in ALL as compared with normal CD34+ progenitor cells (P⩽0.001). Most striking was the 2534-fold higher expression of the recently discovered miR-708 in the B-other group as compared with normal CD34+ cells (P<0.001). In contrast, only 5 out of 19 miRNAs (miR-100, miR-125b, miR-99a, miR-196b and miR-let-7e) are downregulated in B-other as compared with normal CD34+ cells (median 8- to 233-fold; P<0.001).
Intriguing is the 560-fold difference in the expression of miR-196b between MLL-rearranged and B-other cases (P<0.001); miR-196b is 233-fold lower expressed in B-other ALL versus normal CD34+ cells (P<0.001), whereas this miRNA is 2.4-fold higher expressed in MLL-rearranged patients (Table 2; P=0.02). In addition to miR-196b, seven other miRNAs are differentially expressed between MLL-rearranged and other precursor B-ALL cases as shown in Figure 2 (0.001⩽P<0.05), for example, miR-708 is 528-fold higher expressed in B-other than in MLL-rearranged ALL cases (P<0.001; Figure 2, Table 2).
Differential expression of miRNAs in MLL-rearranged ALL and non-MLL rearranged precursor B-ALL patients. Box plots (median and interquartile range) show the distribution of fold change of expression in 16 MLL-translocated and 19 B-other precursor B-ALL cases (negative for known genetic abnormalities) compared with CD34+ samples. MLL-rearranged ALL versus B-other: *P<0.05; **P<0.01 and ***P<0.001 (MWU).
Only one out of five miRNAs tested for its potential linkage to the MLL-chromosomal region (11q23) and/or MLL1 function, that is, miR-34b, showed a ∼2-fold downregulation in MLL-rearranged ALL compared as with B-other ALL cases (P=0.02; Figure 2, Table 2).
Except for miR-196b, none of the differentially expressed miRNAs are located within regions of copy number variation identified by Mullighan et al.30 However, as this copy number variation, that is a deletion of 7p, occurs in a minority of B-other patients (4%), it is unlikely that this deletion can explain the difference in miR-196b expression between B-other and MLL-translocated ALL patients.
Subtype-specific miRNA expression profiles in ALL
As comparison between MLL-rearranged ALL and B-other ALL (negative for all major genetic abnormalities in ALL) revealed striking differences in miR-196b and miR-708 expressions, we wondered whether this reflected a difference in maturation status between both subtypes or was indicative for the leukemia subtype itself. To address this question, miR-196b and miR-708 expression levels were determined in other ALL subtypes, that is T-ALL (n=15) and precursor B-ALL patients positive for TEL-AML1 (n=10), BCR-ABL (n=10) or E2A-PBX (n=8) or with hyperdiploidy (n=10). MiR-196b was median 500-fold higher expressed in MLL-rearranged ALL compared with all other precursor B-ALL cases (P<0.001) and differed 350-fold up to 600-fold between MLL-rearranged ALL and the different subtypes of precursor B-ALL separately (Figure 3a; 0.001⩽P<0.05). As the differentiation status of MLL-positive ALL is less mature than MLL-negative precursor B subtypes, we hypothesized that miR-196b expression levels may reflect the maturation status rather than the genetic subtype. However, analysis of immunoglobulin and T-cell receptor rearrangement patterns as measured for B-cell maturation status24, 31 revealed that 10 out of 16 MLL-rearranged samples with a high miR-196b expression have a mature immunogenotype, and moreover, that one out of four MLL-rearranged samples with a low miR-196b expression has an immature immunogenotype. In addition, the high miR-196b expression level is not restricted to the proB immunophenotype as shown in Supplementary Table S4, and the expression level of miR-196b seems independent of the specific MLL gene rearrangement (Supplementary Table S5). Interestingly, 5 out of 15 T-ALL cases also show 800-fold higher miR-196b expression than the expression of miR-196b in non-MLL-precursor B-ALL cases (Figure 3a; P<0.001). However, only two out of these five cases have an immature phenotype based on CD1 and surface-membrane-bound CD3 expressions (Figure 3a), and the remaining cases are intermediate and mature T-ALL cases. Figure 3b shows that miR-708 is 250- to 6500-fold higher expressed in TEL-AML1, BCR-ABL, E2A-PBX1, hyperdiploid and B-other cases compared with MLL-rearranged and T-ALL patients (P⩽0.01). miR-708 is also differentially expressed among different non-MLL rearranged cases. For example, E2A-PBX1+ and TEL-AML1+ patients show an 8-to 9-fold higher miR-708 expression than BCR-ABL-translocated patients (P<0.01). Similar to miR-196b, the expression level of miR-708 does not reflect the differentiation status of MLL-rearranged and T-ALL cases (Supplementary Table S6). Taken together, our present data do not suggest that high expression of miR-196b or low expression of miR-708 is indicative of a less-differentiated (immature) lymphoid cell type.
Subtype-specific miRNA expression profiles in pediatric ALL. The expression levels of miR-196b (a) and miR-708 (b) were measured in leukemic cells of 92 patients, reflecting different subtypes of pediatric ALL as indicated at the X axis. B-other cases are negative for MLL, TEL/AML1, BCR-ABL, E2A-PBX and hyperdiploidy. The fold change of expression in individual leukemic samples compared with CD34+ samples is shown. In addition, the maturation status of samples is indicated by a black square (immature), black triangle (intermediate) and open square (mature) as defined by Ig/TCR and CD marker expressions (see Materials and methods). Samples without information about the Ig/TCR status are indicated by dots. Horizontal lines indicate the median fold change. MLL-rearranged ALL versus indicated ALL subtypes: *P<0.05; **P<0.01 and ***P<0.001 (MWU).
MicroRNA genes belonging to the same cluster have a similar expression pattern
miRNA genes such as miR-181a and miR-181b belong to the same cluster based on their close location of less than 200 nucleotides apart from each other on chromosome 9q33.3. (181a cluster). Likewise, miR-181c and miR-181d are located next to each other on 19p13 (181c cluster). These grouped genes are thought to be derived from a common ancestor during evolution and are generally co-transcribed.32, 33 In correspondence with this finding, miR-181a and miR-181b as well as miR-181c and miR-181d show a similar relative expression level compared with CD34+ cells in MLL-rearranged, TEL-AML+, BCR-ABL+, E2A-PBX+, hyperdiploid, B-other and T-ALL patients (Figure 4 and Supplementary Table S6, P<0.002). Another cluster of genes (miR-99a cluster on 21q21.1) is represented here by miR-125b-2 and miR-99a. These miRNAs are both lower expressed in MLL-translocated ALL and B-other ALL than in normal CD34+ cells (P<0.001, Supplementary Table S7).
Clustered miRNA genes have a similar expression pattern in ALL subtypes. The expression levels of miR-181a (a), miR-181b (b), miR-181c (c) and miR-181d (d) were measured in leukemic cells of T-ALL (n=10) and precursor B-ALL patients positive for TEL/AML1 (n=10), BCR-ABL (n=10), E2A-PBX (n=8) or hyperdiploidy (HD, n=10). Dots point to the fold change of expression in leukemic samples compared with normal CD34+ samples. MWU test determined significant expression in all ALL subtypes vs normal CD34+ cells with P<0.001 for all miR-181 members. The expression levels did not significantly differ between subtypes (for example, MLL-rearranged vs TEL/AML1-precursor B-cases, P>0.05).
Discussion
By direct cloning of miRNAs expressed in leukemic cells of MLL-rearranged and B-other (precursor B-ALL phenotype without the known genetic abnormalities MLL, TEL-AML1, BCR-ABL, E2A-PBX or hyperdiploidy) subtypes, we identified 105 known and 8 new human miRNA genes. These miRNA genes encode 101 known and 8 new mature miRNAs and miRNA* forms. The newly cloned miRNAs are not conserved between humans and mice and are less frequently cloned from ALL samples than yet known mature miRNAs. Recently, Landgraf et al.34 postulated that predicted hairpin precursors with low clone counts and evolutionarily less-conserved sequences are not cell type-specific and may originate from dsRNA. However, using mature miRNA-specific stem-loop RT-PCR, the expression of these new and less-conserved miRNAs was shown in ALL patients, sometimes even at relatively high levels; for example hsa-miR-1975 is 3.5-fold higher expressed in precursor B-ALL than in normal CD34+ cells (Supplementary Table S2). We also identified a low clone count miRNA that, by the time of submission of this paper, was indicated as miR-708.34, 35 Quantitative PCR revealed that miR-708 was differentially expressed in ALL subtypes (varying between 5- and 2500-fold compared with normal CD34+ cells, Table 2). This exemplifies that even low count miRNAs are of clinical interest. In addition, our data suggest that expression analysis of miRNA genes, which are presently known in the Sanger database, results in an underestimation of miRNAs that may be important for leukemia and/or informative for genetic abnormalities underlying different ALL subtypes.
A set of 19 miRNAs was further validated in an extended group of patients using the above-mentioned stem-loop RT-PCR technique. This technique was chosen because current probe-based array techniques are hampered by the fact that only known miRNAs can be tested. The expression level of 18 miRNAs in MLL-rearranged ALL and all 19 miRNAs in B-other ALL differed from the expression level in normal CD34+ progenitor cells. In contrast with earlier reports showing mainly downregulation of miRNAs in cancer including leukemia,19, 36 we found both upregulated (14/19) and downregulated (5/19) expressions of miRNAs in ALL compared with normal CD34+ cells. Altered miRNA expression levels may lead to an inappropriate expression of target oncoproteins or target tumor suppressors, thereby facilitating the development of leukemia. A typical example is found in chronic lymphocytic leukemia where a deletion of the miR-15a/16-1 cluster abolishes the inhibition of the anti-apoptotic Bcl-2 target oncogene and, as a consequence, facilitates proliferation of these cells.37 Overexpression of miR-21 in adult AML may be linked with reduced expression of tumor suppressor genes such as PTEN, a gene that is known to play a role in leukemogenesis.20, 38
The expression of miRNAs may be regulated by epigenetic modifications such as promoter hypermethylation. Interestingly, the leukemic fusion gene AML1/ETO has been reported to promote the hypermethylation (and hence reduced expression levels) of miR-223 in t(8;21)-positive AML.39, 40 We also identified expression differences between different genetic subtypes of precursor B-ALL that are characterized by leukemia-specific fusion genes such as MLL fusion genes in MLL/11q23-rearranged ALL, TEL/AML1 in t(12;21)-positive ALL and BCR/ABL in t(9;22)-positive ALL (see Table 2; Figure 3). As miRNAs are known to be differentially regulated during development,41, 42, 43, 44 subtype-specific expression of miRNAs such as miR-196b (high in MLL-rearranged ALL) and miR-708 (high in other precursor B-subtypes) might reflect a difference in the differentiation status of leukemic cells; for example, MLL-rearranged ALL is linked to a more immature proB-type compared with a more differentiated common/preB subtype often found in non-MLL precursor B-ALL subtypes.24, 45 However, no linkage between miR-196b and miR-708 expressions and the maturation status of MLL and T-ALL cases were detected in the present study. This observation suggests that the expression level of both miRNAs reflects the differences between leukemic subtypes and is less likely to be associated with the differentiation status.
Recently, the MLL gene-encoded MLL1 methyltransferase protein was found to extensively bind the miR-142 gene on 17q23.2 resulting in the dysregulation of miR-142 expression.29 However, miR-142 was not found to be differentially expressed between MLL-translocated ALL cases compared with other precursor B-ALL cases in the present study. In addition, no aberrant expression levels were found for other miR genes located in the proximity of the MLL gene, including miR-100 and miR-34b. In contrast, a striking difference in the miR-196b expression of 500-fold was found between MLL-rearranged ALL and different non-MLL precursor B-ALL subtypes as well as a 800-fold difference between a subset of T-ALL cases and non-MLL precursor B-ALL cases. The miR-196b gene is located in the HOXA cluster at chromosome 7p15: an area reported to be affected by copy number variation in ⩽5% of non-MLL precursor B-ALL cases and 0% of MLL-rearranged cases.30 (Table 2). Recently, it was suggested that transcriptional activation of this cluster is caused by MLL1 binding and subsequent H3-K4 trimethylation of associated histones.29 As miR-196b is mapped between HOXA9 and HOXA10, the transcriptional activation of HOXA genes by MLL1 might also affect the expression of miR-196b. Indeed, we observed that miR-196b expression correlated with the expression of HOXA9 and HOXA10 in MLL-rearranged cases (data not shown). Overall, these findings point to a possible co-regulation of the HOXA cluster and miR-196b in MLL-translocated ALL. Similar to miRNA–mRNA co-transcriptional regulation, the transcription of miRNA genes can also be co-regulated as described for the miR-17-92 polycistron.21 In the present study, we showed that the family members of the 181a and 181c clusters are also co-expressed at similar levels in pediatric ALL.
Co-regulation of protein-encoding and/or miR-encoding genes may have important regulatory consequences in cell physiology by generating feedback loops that avoid uncontrolled expression of protein-encoding genes (for example, miR-17-92 cluster and E2F1-c-Myc loop).46 Interestingly, the miR-196 family may be involved in the regulation of evolutionarily conserved homeobox (HOX) genes, which are powerful regulators of (lymphoid) development. MiR-196a, which differs only in one nucleotide from miR-196b, has been shown to target the translation of HOXB8 and HOXC8,47 the latter also predicted as a potential target of miR-196b by three computer algorithms that base their prediction on sequence homology, that is miRanda, PicTar and Targetscan (Supplementary Table S8).48, 49, 50 Aberrant miR-196 expression may contribute to leukemogenesis, as dysregulated HOX genes were shown to directly induce leukemia in mice.51 According to the algorithm miRanda, miR-708 also might regulate two target candidate genes linked to leukemia (Supplementary Table S9).48 The first one, IKAROS family zinc finger 4 (IKZF4), associates with its family member IKAROS, a regulator of lymphocyte commitment and differentiation, which contributes to leukemia if dysregulated.52, 53, 54 The second one, the Feline sarcoma (Fes) oncogene, is involved in cell survival and, interestingly, is found to be translocated in acute promyleocytic leukemia.55, 56 Taken together, both miR-708 and miR-196b might inhibit the translation of proteins associated with normal survival and development of lymphocytes. Dysregulation of these proteins by aberrant expression of miR-196b and/or miR-708 might therefore contribute to leukemogenesis. However, additional biological studies have to reveal whether these miRNAs effectively contribute to leukemogenesis or whether the observed upregulation in the case of miR-196b is just a bystander effect of the activated HOXA cluster in MLL-rearranged cases.
In summary, this study revealed new miRNAs that are, in contrast to presently known miRNAs, not evolutionarily conserved. We show that the expression of both known and newly identified miRNA genes varies in genetically and prognostically different subtypes of pediatric ALL and normal CD34+ progenitor cells. At present, our data suggest that the differential expression of miRNAs, such as miR-196b and miR-708, is more associated with the leukemic subtype than with the maturation status of cells. This phenomenon warrants further functional studies of the role of miRNAs in leukemia.
References
Bartel DP . MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004; 116: 281–297.
Ambros V . MicroRNA pathways in flies and worms: growth, death, fat, stress, and timing. Cell 2003; 113: 673–676.
Chen CZ . MicroRNAs as oncogenes and tumor suppressors. N Engl J Med 2005; 353: 1768–1771.
Available at http://www.sanger.ac.uk/Software/Rfam
Lee Y, Jeon K, Lee JT, Kim S, Kim VN . MicroRNA maturation: stepwise processing and subcellular localization. EMBO J 2002; 21: 4663–4670.
Bernstein E, Caudy AA, Hammond SM, Hannon GJ . Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 2001; 409: 363–366.
Hutvagner G, McLachlan J, Pasquinelli AE, Balint E, Tuschl T, Zamore PD . A cellular function for the RNA-interference enzyme Dicer in the maturation of the let-7 small temporal RNA. Science 2001; 293: 834–838.
Ketting RF, Fischer SE, Bernstein E, Sijen T, Hannon GJ, Plasterk RH . Dicer functions in RNA interference and in synthesis of small RNA involved in developmental timing in C. Elegans Genes Dev 2001; 15: 2654–2659.
Grishok A, Pasquinelli AE, Conte D, Li N, Parrish S, Ha I et al. Genes and mechanisms related to RNA interference regulate expression of the small temporal RNAs that control C elegans developmental timing cell. Genes Dev 2001; 106: 23–34.
Meister G . miRNAs get an early start on translational silencing. Cell 2007; 131: 25–28.
Kim VN . MicroRNA biogenesis: coordinated cropping and dicing. Nat Rev Mol Cell Biol 2005; 6: 376–385.
He L, Hannon GJ . MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet 2004; 5: 522–531.
Chen K, Rajewsky N . The evolution of gene regulation by transcription factors and microRNAs. Nat Rev Genet 2007; 8: 93–103.
Olsen PH, Ambros V . The lin-4 regulatory RNA controls developmental timing in Caenorhabditis elegans by blocking LIN-14 protein synthesis after the initiation of translation. Dev Biol 1999; 216: 671–680.
Ambros V . The functions of animal microRNAs. Nature 2004; 431: 350–355.
Takamizawa J, Konishi H, Yanagisawa K, Tomida S, Osada H, Endoh H et al. Reduced expression of the let-7 microRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res 2004; 64: 3753–3756.
Cummins JM, He Y, Leary RJ, Pagliarini R, Diaz Jr LA, Sjoblom T et al The colorectal microRNAome. Proc Natl Acad Sci USA 2006; 103: 3687–3692.
Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E et al. Frequent deletions and down-regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci USA 2002; 99: 15524–15529.
Garzon R, Volinia S, Liu CG, Fernandez-Cymering C, Palumbo T, Pichiorri F et al. MicroRNA signatures associated with cytogenetics and prognosis in acute myeloid leukemia. Blood 2008; 111: 3183–3189.
Jongen-Lavrencic M, Sun SM, Dijkstra MK, Valk PJ, Lowenberg B . MicroRNA expression profiling in relation to the genetic heterogeneity of acute myeloid leukemia. Blood 2008; 111: 5078–5085.
He L, Thomson JM, Hemann MT, Hernando-Monge E, Mu D, Goodson S et al. A microRNA polycistron as a potential human oncogene. Nature 2005; 435: 828–833.
Pui CH, Evans WE . Treatment of acute lymphoblastic leukemia. N Engl J Med 2006; 354: 166–178.
Stam RW, den Boer ML, Schneider P, Nollau P, Horstmann M, Beverloo HB et al. Targeting FLT3 in primary MLL-gene-rearranged infant acute lymphoblastic leukemia. Blood 2005; 106: 2484–2490.
Jansen MW, Corral L, van der Velden VH, Panzer-Grumayer R, Schrappe M, Schrauder A et al. Immunobiological diversity in infant acute lymphoblastic leukemia is related to the occurrence and type of MLL gene rearrangement. Leukemia 2007; 21: 633–641.
Lau NC, Lim LP, Weinstein EG, Bartel DP . An abundant class of tiny RNAs with probable regulatory roles in Caenorhabditis elegans. Science 2001; 294: 858–862.
Available at http://www.tbi.univie.ac.at/~ivo/RNA
Chen C, Ridzon DA, Broomer AJ, Zhou Z, Lee DH, Nguyen JT et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res 2005; 33: e179.
Lim LP, Lau NC, Weinstein EG, Abdelhakim A, Yekta S, Rhoades MW et al. The microRNAs of Caenorhabditis elegans. Genes Dev 2003; 17: 991–1008.
Guenther MG, Jenner RG, Chevalier B, Nakamura T, Croce CM, Canaani E et al. Global and Hox-specific roles for the MLL1 methyltransferase. Proc Natl Acad Sci USA 2005; 102: 8603–8608.
Mullighan CG, Goorha S, Radtke I, Miller CB, Coustan-Smith E, Dalton JD et al. Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia. Nature 2007; 446: 758–764.
van Zelm MC, van der Burg M, de Ridder D, Barendregt BH, de Haas EF, Reinders MJ et al. Ig gene rearrangement steps are initiated in early human precursor B cell subsets and correlate with specific transcription factor expression. J Immunol 2005; 175: 5912–5922.
Yu J, Wang F, Yang GH, Wang FL, Ma YN, Du ZW et al. Human microRNA clusters: genomic organization and expression profile in leukemia cell lines. Biochem Biophys Res Commun 2006; 349: 59–68.
Baskerville S, Bartel DP . Microarray profiling of microRNAs reveals frequent coexpression with neighboring miRNAs and host genes. RNA 2005; 11: 241–247.
Landgraf P, Rusu M, Sheridan R, Sewer A, Iovino N, Aravin A et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 2007; 129: 1401–1414.
Lui WO, Pourmand N, Patterson BK, Fire A . Patterns of known and novel small RNAs in human cervical cancer. Cancer Res 2007; 67: 6031–6043.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D et al. MicroRNA expression profiles classify human cancers. Nature 2005; 435: 834–838.
Cimmino A, Calin GA, Fabbri M, Iorio MV, Ferracin M, Shimizu M et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc Natl Acad Sci USA 2005; 102: 13944–13949.
Yilmaz OH, Valdez R, Theisen BK, Guo W, Ferguson DO, Wu H et al. Pten dependence distinguishes haematopoietic stem cells from leukaemia-initiating cells. Nature 2006; 441: 475–482.
Fazi F, Racanicchi S, Zardo G, Starnes LM, Mancini M, Travaglini L et al. Epigenetic silencing of the myelopoiesis regulator microRNA-223 by the AML1/ETO oncoprotein. Cancer Cell 2007; 12: 457–466.
Nervi C, Fazi F, Grignani F . Oncoproteins, heterochromatin silencing and microRNAs: a new link for leukemogenesis. Epigenetics 2008; 3: 1–4.
Aboobaker AA, Tomancak P, Patel N, Rubin GM, Lai EC . Drosophila microRNAs exhibit diverse spatial expression patterns during embryonic development. Proc Natl Acad Sci USA 2005; 102: 18017–18022.
Ason B, Darnell DK, Wittbrodt B, Berezikov E, Kloosterman WP, Wittbrodt J et al. Differences in vertebrate microRNA expression. Proc Natl Acad Sci USA 2006; 103: 14385–14389.
Kloosterman WP, Wienholds E, de Bruijn E, Kauppinen S, Plasterk RH . In situ detection of miRNAs in animal embryos using LNA-modified oligonucleotide probes. Nat Methods 2006; 3: 27–29.
Wienholds E, Kloosterman WP, Miska E, Alvarez-Saavedra E, Berezikov E, de Bruijn E et al. MicroRNA expression in zebrafish embryonic development. Science 2005; 309: 310–311.
Pieters R, den Boer ML, Durian M, Janka G, Schmiegelow K, Kaspers GJ et al. Relation between age, immunophenotype and in vitro drug resistance in 395 children with acute lymphoblastic leukemia—implications for treatment of infants. Leukemia 1998; 12: 1344–1348.
O’Donnell KA, Wentzel EA, Zeller KI, Dang CV, Mendell JT . c-Myc-regulated microRNAs modulate E2F1 expression. Nature 2005; 435: 839–843.
Yekta S, Shih IH, Bartel DP . MicroRNA-directed cleavage of HOXB8 mRNA. Science 2004; 304: 594–596.
Available at www.microRNA.org.
Available at http://pictar.bio.nyu.edu.
Available at www.targetscan.org.
Buske C, Humphries RK . Homeobox genes in leukemogenesis. Int J Hematol 2000; 71: 301–308.
Molnar A, Georgopoulos K . The Ikaros gene encodes a family of functionally diverse zinc finger DNA-binding proteins. Mol Cell Biol 1994; 14: 8292–8303.
Perdomo J, Holmes M, Chong B, Crossley M . Eos and pegasus, two members of the Ikaros family of proteins with distinct DNA binding activities. J Biol Chem 2000; 275: 38347–38354.
Winandy S, Wu P, Georgopoulos K . A dominant mutation in the Ikaros gene leads to rapid development of leukemia and lymphoma. Cell 1995; 83: 289–299.
Dalla-Favera R, Bregni M, Erikson J, Patterson D, Gallo RC, Croce CM . Human c-myc onc gene is located on the region of chromosome 8 that is translocated in Burkitt lymphoma cells. Proc Natl Acad Sci USA 1982; 79: 7824–7827.
Sheer D, Hiorns LR, Stanley KF, Goodfellow PN, Swallow DM, Povey S et al. Genetic analysis of the 15;17 chromosome translocation associated with acute promyelocytic leukemia. Proc Natl Acad Sci USA 1983; 80: 5007–5011.
Szczepanski T, van der Velden VH, van Dongen JJ . Classification systems for acute and chronic leukaemias. Best Pract Res Clin Haematol 2003; 16: 561–582.
Acknowledgements
We highly appreciate the contribution of CZ Chen (Department of Microbiology and Immunology, Stanford University, USA) for sharing knowledge and technologies to clone miRNAs, for performing computational analyses of miRNA structures as well as for discussing study results. We also thank the members of the COALL study group (headed by GE Janka-Schaub, Hamburg, Germany) and the Interfant study group (headed by R Pieters, Erasmus MC, Rotterdam, NL) for supporting this study with patient samples. We are grateful to MWJC Jansen (Department of Immunology, Erasmus MC, Rotterdam, NL) for providing the maturation status of MLL-rearranged cases. This study was financially supported by the Dutch Cancer Society (Program Grant EUR 2005-3662; MLdB/RP), The Netherlands Organization for Scientific Research (NWO-Vidi Grant; MLdB) and the Baxter foundation (Chen CZ).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Leukemia website (http://www.nature.com/leu)
Supplementary information
Rights and permissions
About this article
Cite this article
Schotte, D., Chau, J., Sylvester, G. et al. Identification of new microRNA genes and aberrant microRNA profiles in childhood acute lymphoblastic leukemia. Leukemia 23, 313–322 (2009). https://doi.org/10.1038/leu.2008.286
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2008.286
Keywords
This article is cited by
-
The advances of E2A-PBX1 fusion in B-cell acute lymphoblastic Leukaemia
Annals of Hematology (2023)
-
miR-1975 serves as an indicator of clinical severity upon influenza infection
European Journal of Clinical Microbiology & Infectious Diseases (2021)
-
CCN2 (Cellular Communication Network factor 2) in the bone marrow microenvironment, normal and malignant hematopoiesis
Journal of Cell Communication and Signaling (2021)
-
MicroRNA-708 is a novel regulator of the Hoxa9 program in myeloid cells
Leukemia (2020)
-
Circulating microRNAs as minimal residual disease biomarkers in childhood acute lymphoblastic leukemia
Journal of Translational Medicine (2019)