2019-10-25
Inferring Disease-Associated MicroRNAs Using Semi-supervised Multi-Label Graph Convolutional Networks
Publication
Publication
iScience , Volume 20 p. 265- 277
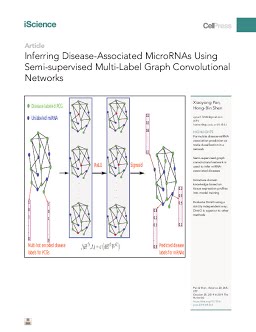
Disease; Gene Network; Biocomputational Method; Computer ModelingMicroRNAs (miRNAs) play crucial roles in biological processes involved in diseases. The associations between diseases and protein-coding genes (PCGs) have been well investigated, and miRNAs interact with PCGs to trigger them to be functional. We present a computational method, DimiG, to infer miRNA-associated diseases using a semi-supervised Graph Convolutional Network model (GCN). DimiG uses a multi-label framework to integrate PCG-PCG interactions, PCG-miRNA interactions, PCG-disease associations, and tissue expression profiles. DimiG is trained on disease-PCG associations and an interaction network using a GCN, which is further used to score associations between diseases and miRNAs. We evaluate DimiG on a benchmark set from verified disease-miRNA associations. Our results demonstrate that DimiG outperforms the best unsupervised method and is comparable to two supervised methods. Three case studies of prostate cancer, lung cancer, and inflammatory bowel disease further demonstrate the efficacy of DimiG, where top miRNAs predicted by DimiG are supported by literature.
| Additional Metadata | |
|---|---|
| , , , | |
| doi.org/10.1016/j.isci.2019.09.013, hdl.handle.net/1765/120180 | |
| iScience | |
| Organisation | Department of Medical Informatics |
|
Pan, X., & Shen, H.-B. (2019). Inferring Disease-Associated MicroRNAs Using Semi-supervised Multi-Label Graph Convolutional Networks. iScience, 20, 265–277. doi:10.1016/j.isci.2019.09.013 |
|