2015
A Testis-Specific Chaperone and the Chromatin Remodeler ISWI Mediate Repackaging of the Paternal Genome
Publication
Publication
Cell Reports , Volume 13 - Issue 7 p. 1310- 1318
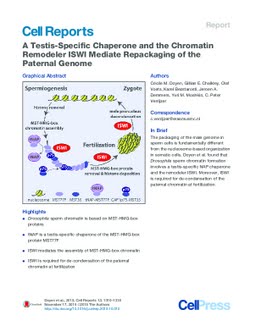
During spermatogenesis, the paternal genome is repackaged into a non-nucleosomal, highly compacted chromatin structure. Bioinformatic analysis revealed that Drosophila sperm chromatin proteins are characterized by a motif related to the high-mobility group (HMG) box, which we termed male-specific transcript (MST)-HMG box. MST77F is a MST-HMG-box protein that forms an essential component of sperm chromatin. The deposition of MST77F onto the paternal genome requires the chaperone function of tNAP, a testis-specific NAP protein. MST77F, in turn, enables the stable incorporation of MST35Ba and MST35Bb into sperm chromatin. Following MST-HMG-box protein deposition, the ATP-dependent chromatin remodeler ISWI mediates the appropriate organization of sperm chromatin. Conversely, at fertilization, maternal ISWI targets the paternal genome and drives its repackaging into de-condensed nucleosomal chromatin. Failure of this transition in ISWI mutant embryos is followed by mitotic defects, aneuploidy, and haploid embryonic divisions. Thus, ISWI enables bi-directional transitions between two fundamentally different forms of chromatin.
| Additional Metadata | |
|---|---|
| doi.org/10.1016/j.celrep.2015.10.010, hdl.handle.net/1765/79248 | |
| Cell Reports | |
|
Doyen, C., Chalkley, G., Voets, O., Bezstarosti, K., Demmers, J., Moshkin, Y., & Verrijzer, P. (2015). A Testis-Specific Chaperone and the Chromatin Remodeler ISWI Mediate Repackaging of the Paternal Genome. Cell Reports, 13(7), 1310–1318. doi:10.1016/j.celrep.2015.10.010 |
|