2016-05-03
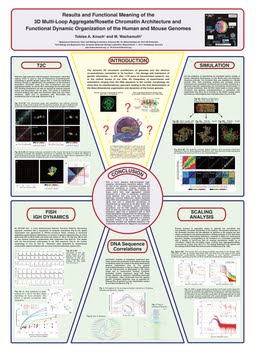
Results and functional meaning of the detailed 3D multi-loop aggregate/rosette chromatin architecture and functional dynamic organization of the human and mouse genomes.
Publication
Publication
The three-dimensional organization of chromosomes of eukaryotic interphase cells is emerging as an important parameter for the regulation of genes as well as the storage, replication and expression of genetic information in general. Whereas a couple of techniques like electrone microscopy (EM), fluorescentce in situ hybridization, or chromosome capture techniques as 3C, 4C, 5C, HiC, or the novel T2C have been used for researching genomic architecture, its dynamics has remained underexplored. Here, we present a novel approach to dissect intramolecular polymer dynamics from fluorescence intensity fluctuations measured with fluorescence correlation spectroscopy (FCS) to investigate the higher order chromatin dynamics in living cells. Using fluorescently tagged linker histone H1 and core histones H2A and H2B as tracer molecules, we found distinct chromatin relaxation times of ~160 ms for open and ~90 ms for dense chromatin areas, corresponding to radii of gyration of 240 and 300 nm for the topologically independent chromatin units. According to their genomic content of ~1 Mb, these domains correspond to subchromosomal domains, already seen by FISH, and histone GFP labelling beforehand and correspond to so called distinct topologically associating domains (TADs). This is an impressive advancements in the field, since details on the dynamic and structural properties of subchromosomal domains/TADs in living cells are scarce. Based on these results, quantitative analytical and numerical modelling provided access to mass density, persistence length and topological information of chromatin. It allowed to extract these parameters from 3C/5C/HiC/T2C results, to predict chromatin conformation and distance data, and to identify complex looping as crucial for domain formation. Especially, in combination with T2C, a recently developed selective high-throughput high-resolution chromosomal interaction capture technique (see abstract T.A. Knoch & M. Wachsmuth, Determination of the three-dimensional organization of chromatin by modelling-supported selective chromosomal interaction capture (T2C)), which provides very good signal-to-noise ratio, we present a comprehensive systematic approach for the understanding of chromatin dynamics and structure as well as for insight into their impact on gene regulation. As an outlook, we show light-sheet microscopy-based FCS maps of chromatin domain dynamics.
| Additional Metadata | |
|---|---|
| , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , , | |
| hdl.handle.net/1765/95863 | |
| Nuclear Organization and Function, Cold Spring Harbor Laboratory, Cold Spring Harbor, New York, USA, 3rd – 7th May, 2016. | |
| Organisation | Biophysical Genomics, Department Cell Biology & Genetics |
|
Knoch, T., & Wachsmuth, M. (2016). Results and functional meaning of the detailed 3D multi-loop aggregate/rosette chromatin architecture and functional dynamic organization of the human and mouse genomes.. Presented at the Nuclear Organization and Function, Cold Spring Harbor Laboratory, Cold Spring Harbor, New York, USA, 3rd – 7th May, 2016. Retrieved from http://hdl.handle.net/1765/95863 |
|